P4V: Difference between revisions
No edit summary |
|||
| (17 intermediate revisions by 3 users not shown) | |||
| Line 7: | Line 7: | ||
For people, who want to write just a simple script rather than an real applet, the easiest way to get informations from the vasprun.xml file is to use the XMLSystemPM class defined in p4vasp.SystemPM. | For people, who want to write just a simple script rather than an real applet, the easiest way to get informations from the vasprun.xml file is to use the XMLSystemPM class defined in p4vasp.SystemPM. | ||
__TOC__ | |||
== Installation == | |||
Download installation for UBUNTU 20.04: | |||
[http://aliga.iciq.es/wiki/images/6/62/p4vasp.ubuntu_20_04.tgz Installation_for_UBUNTU_20_04] | |||
View pdf Installation for UBUNTU OLD VERSIONS: | |||
[http://aliga.iciq.es/wiki/images/0/01/p4vasp_Ubuntu_Installation.pdf p4vasp_ubuntu_installation] | [http://aliga.iciq.es/wiki/images/0/01/p4vasp_Ubuntu_Installation.pdf p4vasp_ubuntu_installation] | ||
| Line 61: | Line 66: | ||
p4v | p4v | ||
== Determine Velocities AutoCorrelations in Molecular Dynamics == | |||
One of the most powerful tool in P4Vasp is the possibility to calculate the autocorrelation of all the velocities in a system just with a few clicks. For molecular dynamics results, this autocorrelation can be usefull, especially for Infra-Red measurements. Here is how to use it: | |||
* Firstly, the autocorrelation can be obtained only from the vasprun.xml file resulting for a finished calculation. If your molecular dynamics was aborted before reaching its last step, then you have to modify the end of your vasprun.xml file. [[nice_termination|Here is a little tool]] written in Python 2.7 to get a proper termination for files recovered from unfinished run. | |||
* Once the file is loaded, you can look at the evolution of the system in time and also choose the "VAC" option in the "MD" menu. | |||
* Then, the autocorrelation can be calculated for all the atoms simply with the "Read VAC" button. | |||
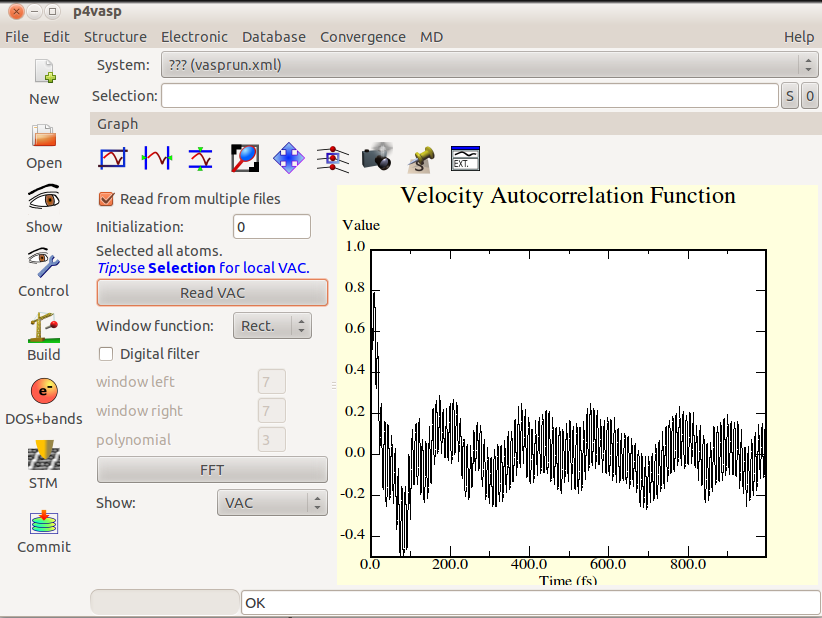
* For a molecular dynamics with N steps, the autocorrelation will be calculated with N-2 positions. Here is a molecular dynamics over 1000 steps. | |||
[[Image:VAC2.png]] | |||
* The Fourier Transformation can be obtained by the "FFT" button and will display useful information about the frequencies existing in the system. | |||
* To get the data in a usable form, the photo camera button will three files, a postscript picture, a dat file with the data and a file with a .agr extension describing the graph. | |||
Latest revision as of 06:30, 26 August 2022
go back to Main Page, Computational Resources, GUIS
P4VASP is a library for processing outputs from a VASP program (http://cms.mpi.univie.ac.at/vasp/Welcome.html) It is focussed mainly on the new xml output format, but it offers also limited support for some older formats (POSCAR, INCAR).
P4Vasp contains also an extensible GUI for visualisation and an api for (reasonably) easy creation of extension applets.
For people, who want to write just a simple script rather than an real applet, the easiest way to get informations from the vasprun.xml file is to use the XMLSystemPM class defined in p4vasp.SystemPM.
Installation[edit]
Download installation for UBUNTU 20.04: Installation_for_UBUNTU_20_04
View pdf Installation for UBUNTU OLD VERSIONS: p4vasp_ubuntu_installation
To install the new version of p4VASP (version 0.3.20) (http://download.wavemol.org/software/p4vasp/p4vasp-0.3.20.tgz) in Ubuntu 10.01, it can see this information (http://cms.mpi.univie.ac.at/vasp-forum/forum_viewtopic.php?2.7812).
Install packages necessary for build:
sudo apt-get install libfltk1.1-dev python2.6-dev g++ libglu1-mesa-dev python-gtk2-dev python-pysqlite2
Extract the p4vasp-0.3.20.tgz in /opt:
mv *p4vasp-0.3.20.tgz /opt cd /opt tar -zxvf p4vasp-0.3.20.tgz cd p4vasp-0.3.20
Optional: if you want to rebuild src/cp4vasp_wrap.cpp:
sudo apt-get install swig rm src/cp4vasp_wrap.cpp
Change Makefile as follows:
vi Makefile
line #19: SITE_PACKAGES = $(PYTHON_HOME)/dist-packages line #143: #chmod -R 755 $(P4VASP_HOME)/doc/api/c
Build and install p4VASP:
make sudo make install
To prevent a Segmentation Fault when running p4VASP:
mv /usr/bin/p4v /usr/lib/p4vasp/p4v.py
and create a new /usr/bin/p4v as follows:
sudo vi /usr/bin/p4v
#!/bin/sh export LD_PRELOAD=libstdc++.so.6 exec /usr/lib/p4vasp/p4v.py
Install packages necessary to run:
sudo apt-get install python-numeric python-numeric-ext
Run p4VASP:
p4v
Determine Velocities AutoCorrelations in Molecular Dynamics[edit]
One of the most powerful tool in P4Vasp is the possibility to calculate the autocorrelation of all the velocities in a system just with a few clicks. For molecular dynamics results, this autocorrelation can be usefull, especially for Infra-Red measurements. Here is how to use it:
- Firstly, the autocorrelation can be obtained only from the vasprun.xml file resulting for a finished calculation. If your molecular dynamics was aborted before reaching its last step, then you have to modify the end of your vasprun.xml file. Here is a little tool written in Python 2.7 to get a proper termination for files recovered from unfinished run.
- Once the file is loaded, you can look at the evolution of the system in time and also choose the "VAC" option in the "MD" menu.
- Then, the autocorrelation can be calculated for all the atoms simply with the "Read VAC" button.
- For a molecular dynamics with N steps, the autocorrelation will be calculated with N-2 positions. Here is a molecular dynamics over 1000 steps.
- The Fourier Transformation can be obtained by the "FFT" button and will display useful information about the frequencies existing in the system.
- To get the data in a usable form, the photo camera button will three files, a postscript picture, a dat file with the data and a file with a .agr extension describing the graph.