Xps.py: Difference between revisions
No edit summary |
No edit summary |
||
| (5 intermediate revisions by 2 users not shown) | |||
| Line 2: | Line 2: | ||
1. Download this script and move xps.py to your ~/bin folder. | |||
1. Download this script and move xps.py to your ~/bin folder. Link to download: [[Image:Xps.tgz]] | |||
2. chmod u+x ~/bin/xps.py | 2. chmod u+x ~/bin/xps.py | ||
| Line 11: | Line 12: | ||
Note: | Note: | ||
1) By default, this script reads POSCAR as input, if you want to use CONTCAR, there are two options: | 1) By default, this script reads POSCAR as input, if you want to use CONTCAR, there are two options: | ||
i) mv CONTCAR POSCAR | i) mv CONTCAR POSCAR | ||
ii) In line 16 of the script, change f = open("POSCAR", 'r') to f = open("CONTCAR", 'r') | ii) In line 16 of the script, change f = open("POSCAR", 'r') to f = open("CONTCAR", 'r') | ||
2) You will get a kind Warning about the INCAR setting in your next XPS calculations, it is not the problem of the script. | |||
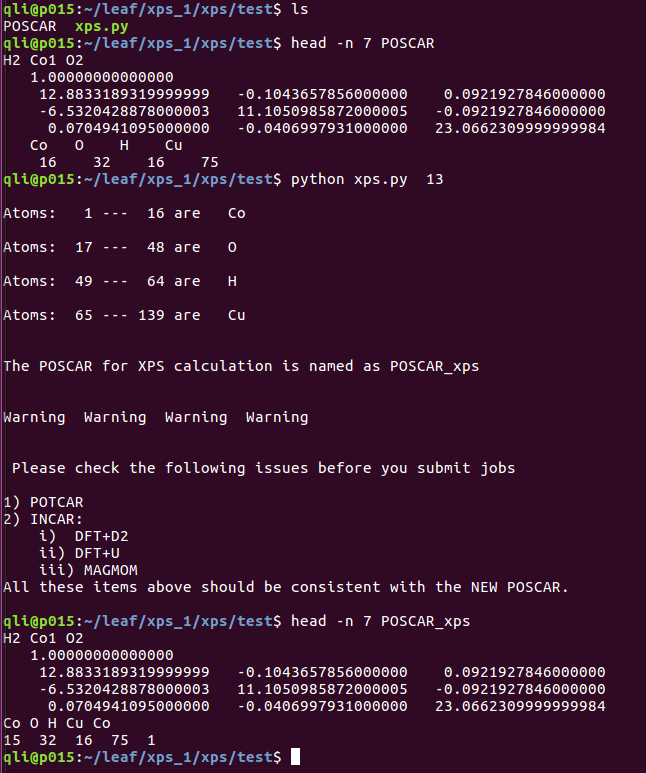
[[Image:Xps.png]] | |||
Enjoy | Enjoy | ||
Latest revision as of 11:30, 8 November 2018
go back to Main Page, Computational Resources, Scripts, Scripts for VASP or VASP
1. Download this script and move xps.py to your ~/bin folder. Link to download: File:Xps.tgz
2. chmod u+x ~/bin/xps.py
3. Go to your calculation directory and run command: xps.py num (num is the number of the atom which you are intersted in)
4 This script will move the selected atom to the end of POSCAR and modify the line 6 and 7 accordingly.
Note:
1) By default, this script reads POSCAR as input, if you want to use CONTCAR, there are two options:
i) mv CONTCAR POSCAR
ii) In line 16 of the script, change f = open("POSCAR", 'r') to f = open("CONTCAR", 'r')
2) You will get a kind Warning about the INCAR setting in your next XPS calculations, it is not the problem of the script.
Enjoy